-Search query
-Search result
Showing 1 - 50 of 122 items for (author: nans & a)

EMDB-18729: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18730: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18731: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18732: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18733: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18734: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed
Method: single particle / : Acton OJ, Sheppard D, Rosenthal PB, Taylor IA

EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion
Method: electron tomography / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M
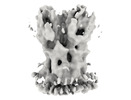
EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15596: 
In situ subtomogram average of Vaccinia virus (WR) palisade, all virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15597: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from CEVs
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15598: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IMVs
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15599: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IEVs
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15600: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (immature virions)
Method: electron tomography / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15601: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (mature virions)
Method: electron tomography / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-15602: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

PDB-8arh: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions
Method: subtomogram averaging / : Calcraft T, Hernandez-Gonzalez M, Nans A, Rosenthal PB, Way M

EMDB-26862: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD
Method: helical / : Li S, Nanson JD, Manik MK, Gu W, Landsberg MJ, Ve T, Kobe B

PDB-7uxu: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD
Method: helical / : Li S, Nanson JD, Manik MK, Gu W, Landsberg MJ, Ve T, Kobe B

EMDB-13978: 
S. cerevisiae CMGE nucleating origin DNA melting
Method: single particle / : Lewis JS, Sousa JS, Costa A

EMDB-13988: 
S. cerevisiae CMGE dimer nucleating origin DNA melting
Method: single particle / : Lewis JS, Sousa JS, Costa A

EMDB-14439: 
S. cerevisiae CMGE dimer nucleating origin DNA melting
Method: single particle / : Lewis JS, Sousa JS, Costa A

PDB-7qhs: 
S. cerevisiae CMGE nucleating origin DNA melting
Method: single particle / : Lewis JS, Sousa JS, Costa A

PDB-7z13: 
S. cerevisiae CMGE dimer nucleating origin DNA melting
Method: single particle / : Lewis JS, Sousa JS, Costa A

EMDB-13439: 
Cryo-EM structure of E. coli TnsB in complex with right end fragment of Tn7 transposon
Method: single particle / : Kaczmarska Z, Czarnocki-Cieciura M, Rawski M, Nowotny M

EMDB-13440: 
Cryo-EM helical reconstruction of E. coli TnsB in complex with right end fragment of Tn7 transposon
Method: helical / : Czarnocki-Cieciura M, Kaczmarska Z

EMDB-26322: 
MVV cleaved synaptic complex (CSC) intasome at 3.4 A resolution
Method: single particle / : Shan Z, Pye VE, Cherepanov P, Lyumkis D

PDB-7u32: 
MVV cleaved synaptic complex (CSC) intasome at 3.4 A resolution
Method: single particle / : Shan Z, Pye VE, Cherepanov P, Lyumkis D

EMDB-26191: 
Cryo-EM structure of human SARM1 TIR domain in complex with 1AD
Method: single particle / : Saikot FK, Kobe B, Ve T, Nanson JD, Gu W, Luo Z, Brillault L, Landsberg MJ

EMDB-14453: 
MVV strand transfer complex (STC) intasome in complex with LEDGF/p75 at 3.5 A resolution
Method: single particle / : Ballandras-Colas A, Nans A, Cherepanov P

EMDB-24272: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-24273: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-24274: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (SAM-TIR:1AD)
Method: single particle / : Kerry PS, Adams S, Cunnea K, Brearley A, Bosanac T, Hughes RO, Ve T

PDB-7nak: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

PDB-7nal: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-13176: 
3.0 A resolution structure of a DNA-loaded MCM double hexamer
Method: single particle / : Greiwe JF, Locke J, Nans A, Costa A

EMDB-13211: 
Structure of a DNA-loaded MCM double hexamer engaged with the Dbf4-dependent kinase
Method: single particle / : Greiwe JF, Locke J, Nans A, Costa A

PDB-7p30: 
3.0 A resolution structure of a DNA-loaded MCM double hexamer
Method: single particle / : Greiwe JF, Miller TCR, Martino F, Costa A

PDB-7p5z: 
Structure of a DNA-loaded MCM double hexamer engaged with the Dbf4-dependent kinase
Method: single particle / : Greiwe JF, Miller TCR, Martino F, Costa A

PDB-7pel: 
CryoEM structure of simian T-cell lymphotropic virus intasome in complex with PP2A regulatory subunit B56 gamma
Method: single particle / : Barski M, Pye VE, Nans A, Cherepanov P, Maertens GN

EMDB-12817: 
Chaetomium thermophilum Chl1 Helicase
Method: single particle / : Hodakova Z, Singleton MR

EMDB-12585: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

EMDB-12586: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (one RBD erect)
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

EMDB-12587: 
Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

PDB-7nt9: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

PDB-7nta: 
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (one RBD erect)
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

PDB-7ntc: 
Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab
Method: single particle / : Rosa A, Pye VE, Nans A, Cherepanov P

EMDB-11928: 
SctV (SsaV) cytoplasmic domain
Method: single particle / : Matthews-Palmer TRS, Gonzalez-Rodriguez N, Calcraft T, Lagercrantz S, Zachs T

PDB-7awa: 
SctV (SsaV) cytoplasmic domain
Method: single particle / : Matthews-Palmer TRS, Gonzalez-Rodriguez N, Calcraft T, Lagercrantz S, Zachs T, Yu XJ, Grabe G, Holden D, Nans A, Rosenthal P, Rouse S, Beeby M
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
